Plant Lipidomics
The Author: Ruth Welti, Kansas Lipidomics Research Center, Division of Biology, Kansas State University, Manhattan, KS 66506, USA
1. Introduction
“Lipidomics” refers to detailed, comprehensive profiling of lipid molecular species, often in relation to genotype. Comparison of the lipid profiles of wild-type plants with those of plants that have been subjected to forward- or reverse-genetic manipulation, in parallel with developmental and physiological phenotyping, can aid in characterization of the roles of the manipulated genes and enzymes [e.g. 1-9]. For example, analysis of lipid changes can provide clues as to the physiological substrates and products of genetically altered enzymes [1].
In plant lipidomic analysis, quantitative information on numerous individual lipid species is typically acquired directly from organic extracts of plant material without chemical modification. Complex lipids, and especially membrane lipids, are particularly amenable to these direct, comprehensive analyses. Information related to plant lipidomics and similar to that in this article has recently been published online in The Arabidopsis Book [10].
Although it is possible to conduct a large-scale lipid analysis by traditional analytical technologies, such as thin-layer chromatography with gas chromatography, or by nuclear magnetic resonance methods, mass spectrometry is the leading technology for large-scale and rapid “lipidomic” analysis. Mass spectrometry-based lipidomics is rapid in comparison to “traditional” lipid analysis and requires relatively small amounts of material (i.e. 0.1 mg of leaf dry weight is sufficient). There are several possible mass spectrometry-based approaches. One approach is to obtain chemical formulas of intact lipid ions formed by ionization in the mass spectrometer by obtaining accurate m/z (mass/charge ratio, where z is typically = 1) by Fourier transform ion cyclotron resonance mass spectrometry [11]. An advantage of this approach is that it is conceptually simple; theoretically an analysis could be derived from a single spectrum. In practice, combining spectra acquired over small m/z ranges increases the sensitivity of this technique [12]. A disadvantage of accurate m/z as a means of lipid identification is that many chemical formulas are shared by several lipid molecular species; thus additional data are required to identify specific compounds.
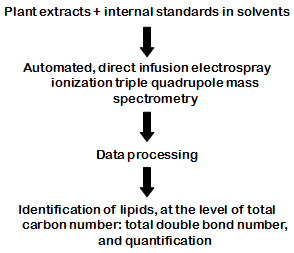
The second and most utilized mass spectrometry-based lipidomic approach employs analysis of pairs of intact lipid ion m/z and fragment m/z as a means of identification and quantification of lipid species. This analysis is efficiently performed using a triple quadrupole mass spectrometer. Plant phospholipids and glycolipids, in particular, are amenable to analysis by direct infusion electrospray ionization triple quadrupole mass spectrometry [overview in Fig. 1].
Figure 1. Direct and rapid analysis of plant polar lipids by electrospray ionization triple quadrupole mass spectrometry, a.k.a. lipidomic analysis.
2. Precursor and Neutral Loss Scanning
The direct infusion electrospray ionization triple quadrupole tandem mass spectrometry approach most applied to plant polar lipids [13] utilizes a series of “precursor” and “neutral loss” scans [based on 14]. This method takes advantage of the formation of common fragments from related complex lipids upon collision-induced dissociation in a triple quadrupole mass spectrometer. A triple quadrupole mass spectrometer has two analyzer quadrupoles (Q1 and Q3) with a collision cell (the second quadrupole, Q2) in between. Q1 passes intact lipid ions to the collision cell, where the ions are fragmented and passed to Q2; Q3 passes fragment ions to the detector.
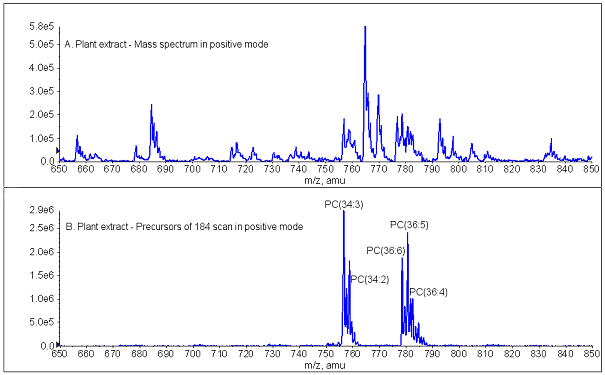
In polar lipids, the analyzed collision-induced dissociation fragment is often a head group fragment common to all members of a lipid class. For example, phosphatidylcholine molecular species, which vary in fatty acid composition, produce a common phosphocholine fragment (C5H15O4PN)+ with a single positive charge and a m/z of 184. Thus, phosphatidylcholine molecular species can be identified by the formation of a m/z 184 fragment plus the precursor (intact) ion fragment m/z. If the common fragment is charged, like the fragment produced by phosphatidylcholine molecules, a scan for the precursors of the fragment (a “precursor scan”) is the analyzed spectrum (plot of signal vs. Q1 m/z) [13]. Figure 2 illustrates the specificity afforded by use of the precursor scan mode. A single MS scan [Fig. 2A] of a directly infused extract depicts total intact ions, arising from multiple lipid classes. Figure 2B shows a precursor scan of the same directly infused extract; this scan depicts the intact ion/fragment pairs delineating lipid species within a class (phosphatidylcholines). When a group/class of lipids fragments such that its common fragment is uncharged, a neutral loss scan (rather than a precursor scan) reveals the m/z values of the lipid molecular species that contain the neutral fragment (i.e. the members of that group/class).
Figure 2. Mass spectra of phosphatidylcholine from the plant Arabidopsis thaliana. Positive ion spectra were acquired during infusion of a total plant lipid extract mixed with internal standards in chloroform-methanol-300 mM ammonium acetate in water (300/665/35). (A) Electrospray ionization single mass spectrum. Many classes of lipids, including phosphatidylcholines, phosphatidylethanolamines, phosphatidylserines, and monogalactosyldiacylglycerols, produce ions detectable in the positive mode. (B) Electrospray ionization triple quadrupole mass spectrum (of the same extract) for “Precursors of m/z 184”, a fragment derived from the phosphatidylcholine head group. The spectrum shows only phosphatidylcholine molecular species. The labels indicate total acyl carbons: total acyl double bonds.
3. Lipid Profiling by Direct Infusion Electrospray Ionization Triple Quadrupole Mass Spectrometry
A lipid profile is assembled by sequentially carrying out a series of precursor and/or neutral loss scans characteristic of the lipid groups or classes to be analyzed while the extract is continuously infused into the mass spectrometer. Essentially, each of these scans allows identification and quantification of the molecular species of one class or group of lipids. Scans for analysis of many complex plant lipid classes are shown in Table 1.
| Table 1. Precursor and neutral loss scans utilizing characteristic fragments generated by electrospray ionization for analysis of polar complex lipids from the plant Arabidopsis thaliana. These scans are recommended for analysis of lipids in chloroform-methanol-300 mM ammonium acetae in water (300:665:35, v/v/v) | ||||
| Lipids analyzed | Polarity | Ion analyzed | Scan mode | References |
| Phospholipids | ||||
| phosphatidylcholines | + | [M + H]+ | Precursors of m/z 184 | [14] |
| phosphatidylethanolamines | + | [M + H]+ | Neutral loss of 141 | [14] |
| phosphatidylserines | + | [M + H]+ | Neutral loss of 185 | [14] |
| phosphatidylglycerols | + | [M + NH4]+ | Neutral loss of 189 | [15] |
| phosphatidylinositols | + | [M + NH4]+ | Neutral loss of 277 | [14] |
| phosphatidic acids | + | [M + NH4]+ | Neutral loss of 115 | |
|
Sphingolipids |
||||
| glycosylinositolphosphoceramides | + | [M + NH4]+ | Neutral loss of 615 or 179 | |
| hexosylceramides | + | [M + H]+ | Neutral loss of 162 | |
|
Galactolipids |
||||
| sulfoquinovosyldiacylglycerols | - | [M - H]- | Precursors of m/z 225 | [16,17] |
| monogalactosyldiacylglycerols | + | [M + NH4]+ | Neutral loss of 179 | [18] |
| digalactosyldiacylglycerols | + | [M + NH4]+ | Neutral loss of 341 | [18,19] |
| trigalactosyldiacylglycerols | + | [M + NH4]+ | Neutral loss of 503 | [18] |
| tetragalactosyldiacylglycerols | + | [M + NH4]+ | Neutral loss of 665 | [18] |
|
Oxidized polar lipids (by acyl-specific scans) |
||||
| lipids containing 18:4-O or 18:3-2O (where O and 2O indicate “extra” oxygens beyond those found in normal fatty acids) | - | [M - H]- | Precursors of m/z 291 | [9] |
| lipids containing 18:3-O or 18:2-2O | - | [M - H]- | Precursors of m/z 293 | [9] |
| lipids containing 18:2-O or 18:1-2O | - | [M - H]- | Precursors of m/z 295 | [9] |
Each precursor and neutral loss scan provides the m/z values and signals of the intact ions within a class or group. Given the class or group classification, the intact ion m/z can be interpreted as the total number of carbons and double bonds in the component acyl (or sphingosine) chains. If more complete characterization of the complex lipid molecule is required, further analysis, such as mass spectral product ion analysis to identify individual fatty acyl components, needs to be performed. Devaiah et al. [4] utilized product ion analysis to characterize many Arabidopsis polar glycerolipid molecular species in terms of fatty acyl composition.
For each detected lipid molecular species, the mass spectral signal size allows quantification. To achieve accurate quantification by direct infusion electrospray ionization triple quadrupole mass spectrometry, a large number of internal standard compounds, optimally at least two non-naturally-occurring compounds, related to the biological compounds in each class or group, are added to the plant samples [e.g. 1,4,14]. The lipid compounds in a biological mixture are quantified by comparison of the signals from the biological compounds with the signals from the internal standards. Alternatively, relative quantification among samples can be achieved by comparing mass spectral responses to an arbitrary standard compound detected with the same polarity (i.e. positive or negative mode) and scanning mode (i.e. precursor or neutral loss scanning) as the lipids of interest (e.g. detection of oxylipin-containing complex lipids [9]).
4. Liquid Chromatography Can Be Added
Thus far, the most comprehensive lipidomics analyses have been performed by direct infusion, which allows scanning for a large number of lipid classes and species in a short time. However, separating lipid components prior to mass spectrometry by liquid chromatography can reduce ion suppression during electrospray ionization and thus increase sensitivity. Liquid chromatography can also resolve compounds, such as isomers, not resolvable by direct infusion. Liquid chromatographic mass spectral analyses of sphingolipids, other polar lipids, free oxylipins, and acyl CoAs have been performed [20-23].
5. Challenges in Lipidomics
Challenges remaining in lipidomics include:
- Standardizing lipidomics protocols
- Increasing availability of appropriate internal standards
- Establishment of analyses for additional groups of lipids
- Development of a standardized data processing system for interpreting mass spectral data to produce lipid profiles
- Development of a web-accessible lipid profile database that facilitates integration with genomic, gene expression, proteomic, and other metabolomic data
- Increasing access to lipidomics technologies
Acknowledgments
The development of plant lipidomics methodology was supported by the National Science Foundation (MCB 0455318 and MCB 0920663) to RW, Xuemin Wang, Jyoti Shah, Todd Williams, and Gary Gadbury. RW is very grateful to these, other colleagues, and lab members who have contributed to plant lipidomics development.
References
- Welti, R., Li, W., Li, M., Sang, Y., Biesiada, H., Zhou, H.-E., Rajashekar, C.B., Williams, T.D. and Wang, X. Profiling membrane lipids in plant stress responses: Role of phospholipase D-α in freezing-induced lipid changes in Arabidopsis. J. Biol. Chem., 277, 31994-32002 (2002).
- Nandi, A., Krothapalli, K., Buseman, C.M., Li, M., Welti, R., Enyedi, A., and Shah, J. The Arabidopsis thaliana sfd mutants affect plastidic lipid composition and suppress dwarfing, cell death and the enhanced disease resistance phenotypes resulting from the deficiency of a fatty acid desaturase. Plant Cell, 15, 2383-2398 (2003).
- Cruz-Ramírez, A., Oropeza-Aburto, A., Razo-Hernandez, F., Ramirez-Chavez, E. and Herrera-Estrella, L. Phospholipase DZ2 plays an important role in extraplastidic galactolipid biosynthesis and phosphate recycling in Arabidopsis roots. Proc. Natl. Acad. Sci. U.S.A., 103, 6765-6770 (2006).
- Devaiah, S.P., Roth, M.R., Baughman, E., Li, M., Tamura, P., Jeannotte, R., Welti, R. and Wang, X. Quantitative profiling of polar glycerolipid species and the role of phospholipase Dα1 in defining the lipid species in Arabidopsis tissues. Phytochemistry, 67, 1907-1924 (2006).
- Li, M., Welti, R. and Wang, X. Quantitative profiling of Arabidopsis polar glycerolipids in response to phosphorus starvation: Roles of PLDζ1 and PLDζ2 in phosphatidylcholine hydrolysis and digalactosyldiacylglycerol accumulation in phosphorus-starved plants. Plant Physiol., 142, 750-761 (2006).
- Chen, M., Markham, J.E., Dietrich, C.R., Jaworski, J.G. and Cahoon, E.B. Sphingolipid long-chain base hydroxylation is important for growth and regulation of sphingolipid content and composition in Arabidopsis. Plant Cell, 20, 1862-1878 (2008).
- Welti, R., Shah, J., Li, W., Li, M., Chen, J., Burke, J.J., Fauconnier, M.-L., Chapman, K., Chye, M.-L. and Wang, X. Plant lipidomics: Discerning biological function by profiling plant complex lipids using mass spectrometry. Frontiers Biosci., 12, 2494-2506 (2007).
- Li, W., Wang, R., Li, M., Li, L., Wang, C., Welti, R. and Wang, X. Differential degradation of extraplastidic and plastidic lipids during freezing and post-freezing recovery in Arabidopsis thaliana. J. Biol. Chem., 283, 461-468 (2008).
- Maeda, H., Sage, T.L., Isaac, G., Welti, R. and DellaPenna, D. Tocopherols modulate extra-plastidic polyunsaturated fatty acid metabolism in Arabidopsis at low temperature. Plant Cell, 20, 452-470 (2008).
- Li-Beisson, Y., Shorrosh, B., Beisson, F., Andersson, M.X., Arondel, V., Bates, P.D., Baud, S., Bird, D., DeBono, A., Durrett, T.P., Franke, R. B., Graham, I.A., Katayama, K., Kelly, A.A., Larson, T., Markham, J.E., Miquel, M., Molina, I., Nishida, I., Rowland, O., Samuels, L., Schmid, K.M., Wada, H., Welti, R., Xu, C., Zallot, R. and Ohlrogge, J. Acyl lipid metabolism: June 11, 2010. The Arabidopsis Book. (American Society of Plant Biologists, Rockville, MD), doi: 10.1199/tab.0133, www.aspb.org/publications/arabidopsis (2010).
- Iijima, Y., Nakamura, Y., Ogata, Y., Tanaka, K., Sakurai, N., Suda, K., Suzuki, T., Suzuki, H.,Okazaki, K., Kitayama, M., Kanaya, S., Aoki, K. and Shibata, D. Metabolite annotations based on the integration of mass spectral information. Plant J., 54, 949-962 (2008).
- Southam, A.D., Payne, T.G., Cooper, H.J., Arvantis, T.N. and Viant, M.R. Dynamic range and mass accuracy of wide-scan direct infusion nanoelectrospray Fourier transform ion cyclotron resonance mass spectrometry-based metabolomics increased by spectral stitching method. Anal. Chem., 79, 4595-4602 (2007).
- Welti, R. and Wang, X. Lipid species profiling: A high throughput approach to identify lipid compositional changes and determine the function of genes involved in lipid metabolism and signaling. Curr. Opin. Plant Biol., 7, 337-344 (2004).
- Brügger, B., Erben, G., Sandhoff, R., Wieland, F.T. and Lehmann, W.D. Quantitative analysis of biological membrane lipids at the low picomole level by nano-electrospray ionization tandem mass spectrometry. Proc. Natl. Acad. Sci. U.S.A., 94, 2339-2344 (1997).
- Taguchi, R., Houjou, T., Nakanishi, H., Yamazaki, T., Ishida, M., Imagawa, M. and Shimizu, T. Focused lipidomics by tandem mass spectrometry. J. Chromatogr. B. Analyt. Technol. Biomed. Life Sci., 823, 26-36 (2005).
- Gage, D.A., Huang, Z.H. and Benning, C. Comparison of sulfoquinovosyl diacylglycerol from spinach and the purple bacterium Rhodobacter sphaeroides by fast atom bombardment tandem mass spectrometry. Lipids, 27, 632-636 (1992).
- Welti, R., Wang, X. and Williams, T.D. Electrospray ionization tandem mass spectrometry scan modes for plant chloroplast lipids. Anal. Biochem., 314, 149-152 (2003).
- Isaac, G., Jeannotte, R., Esch, S.W. and Welti, R. New mass spectrometry-based strategies for lipids. Gen. Eng. Rev., 28, 129-157 (2007).
- Moreau, R.A., Doehlert, D.C., Welti, R., Isaac, G., Roth, M., Tamura, P. and Nuñez, A. The identification of mono-, di-, tri-, and tetragalactosyl-diacylglycerols and their natural estolides in oat kernels. Lipids, 43, 533-548 (2008).
- Markham, J.E. and Jaworski, J.G. Rapid measurement of sphingolipids from Arabidopsis thaliana by reversed-phase high-performance liquid chromatography coupled to electrospray ionization tandem mass spectrometry. Rapid Commun. Mass Spectrom., 21, 1304-1314 (2007).
- Yang, S., Qiao, B., Lu, S.H., and Yuan, Y.J. Comparative lipidomics analysis of cellular development and apoptosis in two Taxus cell lines. Biochim. Biophys. Acta, 1771, 600-612 (2007).
- Pan, X., Welti, R. and Wang, X. Quantitative analysis of major plant hormones in crude plant extracts by high-performance liquid chromatography-mass spectrometry. Nat. Protoc., 5, 986-992 (2010).
- Larson, T.R. and Graham, I.A. Technical advance: A novel technique for the sensitive quantification of acyl CoA esters from plant tissues. Plant J., 25, 115-125 (2001).
In This Section
- Plant Fatty Acid Synthesis
- Production of Unusual Fatty Acids in Plants
- Arabidopsis Acyl-Coenzyme A-Binding Proteins
- Long Chain acyl-coA Synthetases and Other Acyl Activating Enzymes
- Plant Triacylglycerol Synthesis
- Triacylglycerol Biosynthesis in Eukaryotic Microalgae
- Subcellular Oil Droplets and Oleosins in Plants
- Triacylglycerol Mobilisation in Plants
- Role of Transcription Factors in Storage Lipid Accumulation in Plants
- Biosynthesis of Plant Lipid Polyesters
- Rubber Biosynthesis in Plants
- Carotenoid Biosynthesis and Regulation in Plants
- The Oxylipin Biosynthetic Pathways in Plants
- N-Acylphosphatidylethanolamines (NAPEs), N-acylethanolamines (NAEs) and Other Acylamides: Metabolism, Occurrence and Functions in Plants
- Phosphoinositide Signaling in Plants
- Plant Lipidomics
- 50 years of Galactolipid Research: The Beginnings
- Transport and function of lipids in the plant phloem